Structured patterns
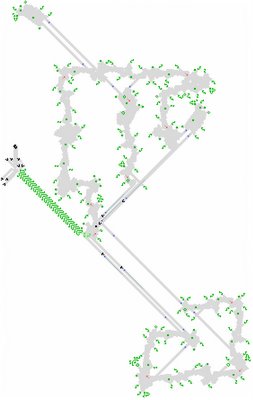
Something I've been wanting to do for a while now is to produce a structured description of Gabriel Nivasch's Caterpillar.In the context of the Game of Life, a "structured pattern" would be a description in terms of the smaller subpatterns that are repeated many times in the full pattern, such as the "shish-ke-babs" or the various rakes that make up the Caterpillar. The subpatterns in turn may be composed of smaller sub-subpatterns like pi climbers, blinkers, gliders, etc. A structured format makes a much better "blueprint" of a large pattern than a standard RLE encoding or picture-format file, and it's also very compact -- probably better compression than any standard binary-archive tool could manage.
Xlife 3.5/3.6 provides a good pattern format for describing this kind of structured pattern. Unfortunately it isn't quite versatile enough for this purpose. The big problem is that when you're dealing with periodic subpatterns, you often want to be able to include copies of them at several different phases. Xlife's approach is to define one phase of each periodic pattern as its "base" pattern; to paste multiple copies in, you paste the first base pattern, run the entire universe some number of generations to get the first pattern to the right phase relative to the second; then paste the second base pattern -- and if necessary run N more generations and paste a third base pattern, and so forth.
For stable patterns, or patterns that don't interact too closely, that's fine. But unfortunately this approach doesn't work very well for large numbers of subpatterns or higher periods -- sometimes Xlife's format doesn't support a "natural" compact definition based on subpatterns, because (for example) two of the subpatterns interfere with each other before the correct phase of the third subpattern can be placed to stabilize the whole conglomeration.
Ideas for an extension of Xlife #I format:
First, for reference, here is a summary of current Xlife syntax.
An extension of the format would fix the problem described above -- you'd have to be able to say, "run just this pattern in isolation for N generations, then paste it at (x,y)." This seems like a more natural way to compose patterns from subpatterns.
-- In fact, that's how Xlife's GUI does it! To register subpatterns in a "load script", you specify subpatterns' locations, transformations, and number of ticks forward in time from the base pattern. It's actually quite a surprise to look in an #I-format file for the first time and find out that Xlife has translated your registered subpatterns into this alternate format that is only sometimes equivalent.
I think Eugene Langvagen's PLife project has the necessary versatility to describe this kind of run-N-ticks-then-paste operation, but I haven't done enough benchmarking yet to know whether PLife can handle Caterpillar-sized structured patterns.
The main problem that I see is that a PLife script can easily become much too syntactically complex to be read directly by other Life software. You could create a structured pattern using a PLife script, but you'd have to convert it back to RLE to communicate it to anyone who doesn't have Python and PLife available. Xlife format is much more straightforward to write a parser for.